Frontiers | Standardization of Sequencing Coverage Depth in NGS: Recommendation for Detection of Clonal and Subclonal Mutations in Cancer Diagnostics

Standardization of Sequencing Coverage Depth in NGS: Recommendation for Detection of Clonal and Subclonal Mutations in Cancer Diagnostics. - Abstract - Europe PMC

A beginner's guide to low‐coverage whole genome sequencing for population genomics - Lou - 2021 - Molecular Ecology - Wiley Online Library

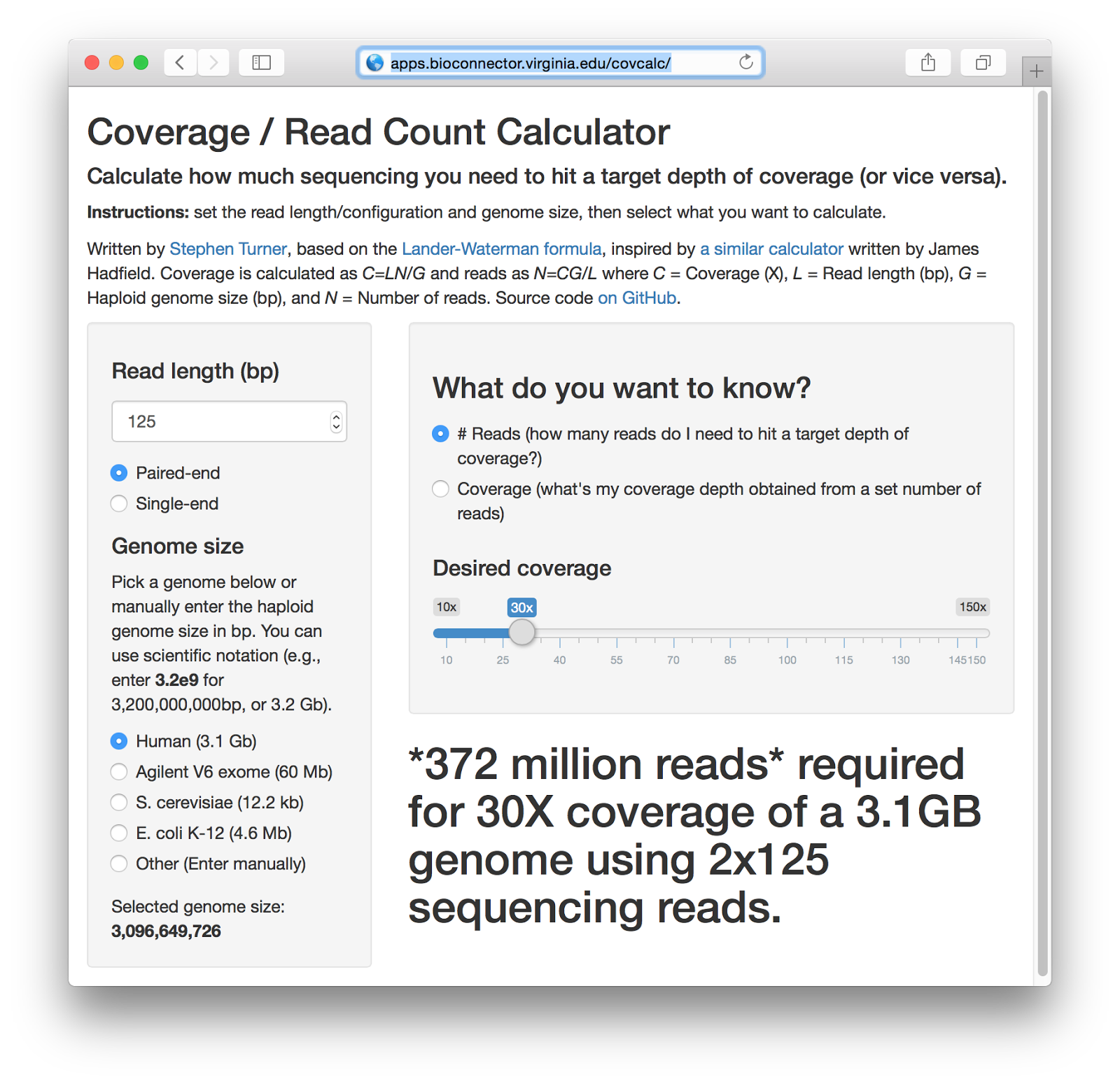
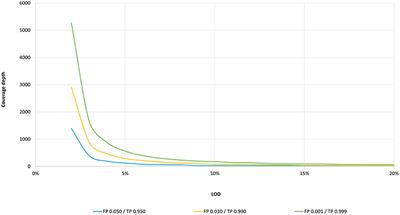
Mathematical Framework Provides a Read Depth Calculator and Guidelines... | Download Scientific Diagram

PDF) Standardization of Sequencing Coverage Depth in NGS: Recommendation for Detection of Clonal and Subclonal Mutations in Cancer Diagnostics